As we kick off a new semester, we’re proud to reflect on the milestones achieved this year in computational medicine. 2023 was a year of learning, growth, and collaboration.
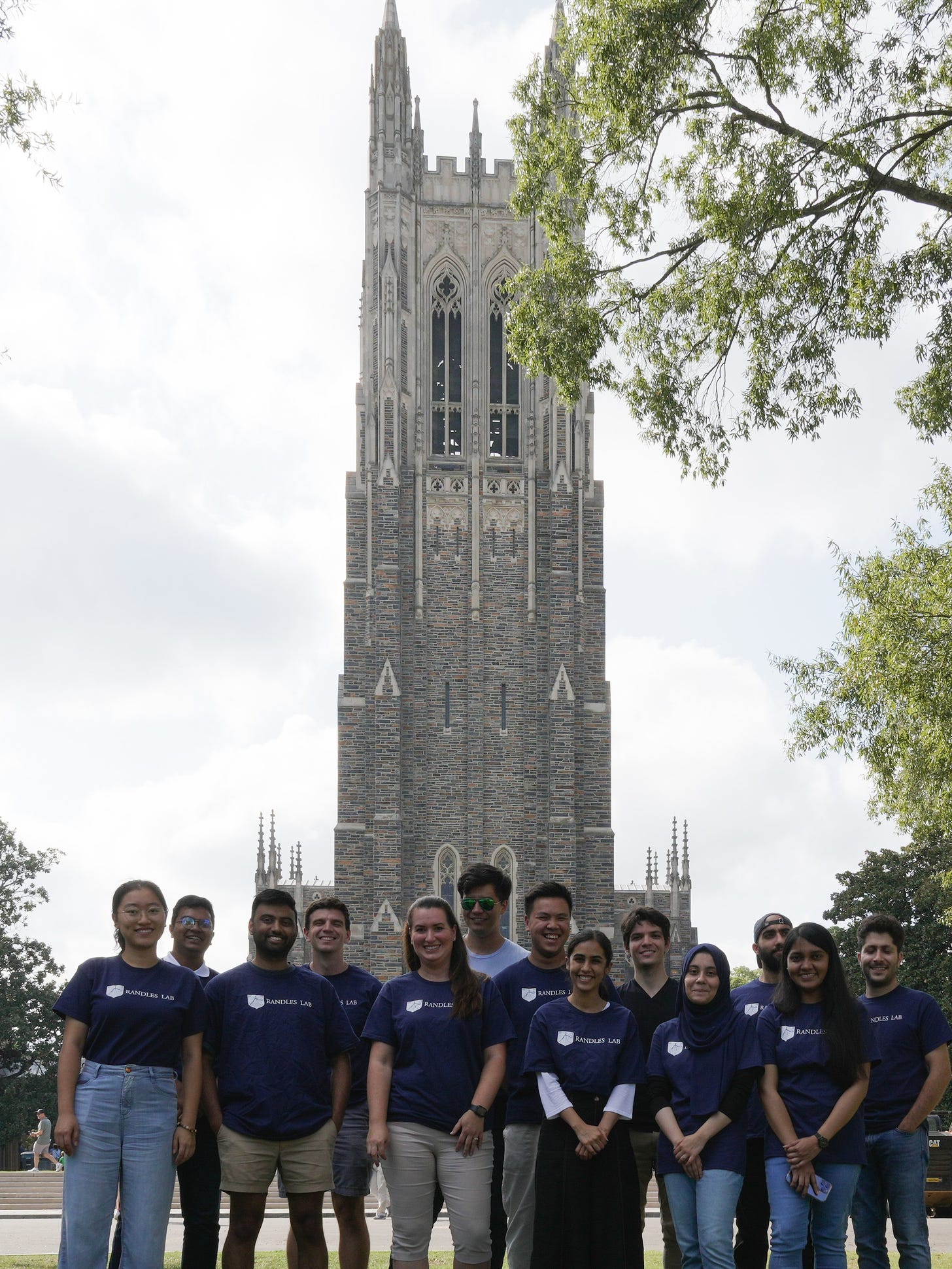
Randles Lab 2023
Academic Contributions
Our presence in the scientific dialogue was marked by 4 journal articles, 7 refereed conference articles, 7 contributed talks, and 9 conference posters. Each publication and presentation allowed us to share our passion for research and contribute to the broader conversation in high performance computing.
Select Highlights from Our 2023 Research:
· Pioneering Longitudinal Hemodynamic Maps: At the 2023 ACM/IEEE International Conference, we introduced a cloud-based framework for personalized 3D hemodynamic maps made possible by advances in wearable technology. Tanade et al. SC23.
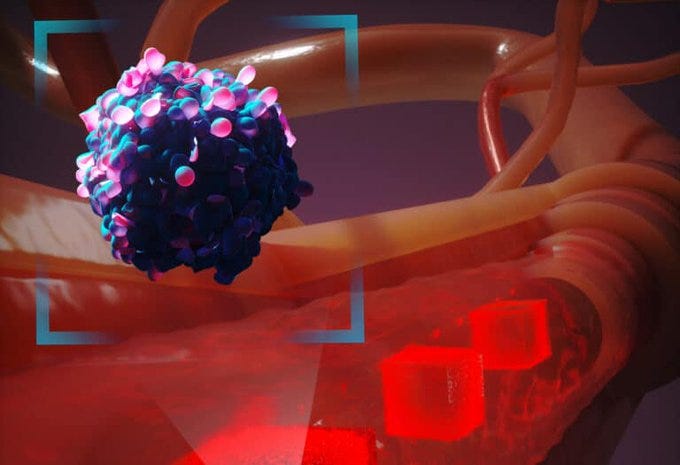
· Refining Cancer Cell Transport Simulations: Our improved APR method now offers more nuanced models of cancer cell movement through bloodstreams, informing potential new paths for treatment. Roychowdhury et al. SC23.
· Assessing different programming models: As we enter the exascale era, systems are becoming increasingly heterogeneous, and the number of programming models is expanding. In our P3HPC paper, we investigated the performance of different programming models for a wide range of GPU-based architectures. Martin et al. P3HPC23.
· Visualizing millions of cells: Handling large-scale fluid-structure-interaction simulations creates a significant data problem. In a series of papers this year, we investigated different in situ and post hoc visualization strategies. Yousef et al. ISAV23. Yousef et al. LDAV23.
· Optimizing use of cloud resources: Cloud computing offers a strong alternative for medical work and applications for users who may not own large, on-premises systems. We developed techniques to integrate cloud computing optimally into different hemodynamic workflows. Ladd et al, IPDPS2023.
· Individualizing ECMO Strategies: Published in Perfusion, our simulations emphasize the critical role of patient-specific factors in the efficacy of ECMO, a life-support technology. Feiger et al. Perfusion, 2023.
· Modeling Tumor Growth Dynamics: We've embraced the challenge of 3D tumor growth simulation, enhancing our ability to model and understand cancer development. Tanade et al. Journal of Computational Science, 2023.
· Quantifying Cellular Configurations: Our work on spatial metrics for cellular distributions is helping to refine simulation accuracy for various physiological processes. Roychowdhury et al. Journal of Computational Science, 2023.
· Assessing the effect of cell heterogeneity: Our work published in Cellular and Molecular Bioengineering revealed that minor variations in the biophysical properties of tumor cells, such as size and membrane elasticity, significantly affect their circulation dynamics, highlighting the critical role of cellular heterogeneity in cancer metastasis research and therapy development.
· Understanding Erythrocyte Deformation: By exploring erythrocyte mechanics, we're contributing to a foundational understanding of how these cells respond to physical stress, with implications for medical devices and therapies. Pepona et al. Computers and Mathematics with Applications, 2023.
Image: Representation of our adaptive mesh refinement (APR) framework that captures cellular interaction over long-length scales.
Recognition and Tenure
The year was also significant for me in particular. I was incredibly honored to be awarded the SIGHPC Emerging Woman Leader in Technical Computing Award and the Duke Stansell Family Distinguished Research Award. In a huge career milestone, I was awarded tenure from Duke University this year and am excited to celebrate it with my lab. They surprised me with a fantastic party with my family included, Zoomed in collaborators, mentors, and lab alums, and made a heartfelt video. I couldn't have achieved it without them, and I am constantly reminded of how lucky I am to be surrounded by such a great team.
Grants, Awards, and Collaborative Ventures
Our research was bolstered by computing awards for leadership-class systems. We continue to prepare for Aurora as part of the Early Science Program, but this year, we were also awarded a DOE INCITE Award and the LLNL Grand Challenge Award. Additionally, we're excited to contribute to the NIH R01 project spearheaded by Lydia Sohn and Mark Labarge and a DOE BRaVE award led by ORNL’s Heidi Hanson.
Knowledge Dissemination and Community Building
Our findings on digital twins and the human circulatory system were disseminated through an invited talk at SC23, a plenary at CompBioMed, and a keynote at IHPCSS. These platforms allowed us to share our work and vision with the global community.
Lab Growth and Milestones
The lab welcomed 3 new postdocs, 2 new graduate students, 1 summer intern, and 1 new research associate. We also celebrated 1 Ph.D. defense, the first Ph.D. hooding in our lab's history, 1 master's graduation, and 3 graduate students passing their prelims—each a stepping stone towards future successes.
Engaging Beyond Research
With the launch of the Computational Medicine Virtual Seminar Series and the Computational Medicine Summer Workshops, we are leading an initiative to cultivate a community in Computational Medicine at Duke. Kicking off the activities and bringing everyone together has been rewarding already.
Computational Medicine Coffee Hour
Community Events and Team Spirit
Hosting the Argonne/Intel hackathon exemplified our commitment to collaborative problem-solving. Our team spirit flourished through various outings and activities, affirming that while we take our research seriously, we value joy and camaraderie just as highly. We’re excited to return to conference travel with team trips to IPDPS, BMES, and SC23. We had a return to our monthly happy hours and lots of fun team activities like a group trip to the Museum of Life and Science, box seats at a Durham Bulls game, and our annual End of Year party.
Team at BMES
Team at SC23
Durham Bulls Game
Looking back, we are grateful for the memories created and the milestones achieved in 2023. As we start 2024, we are excited to see how our efforts in developing digital twins advance and the lab continue to grow and flourish.
Randles Lab
Stay Connected and Explore with Us
Twitter General: @RandlesLab
Twitter Pubs: @RandlesLabPubs
https://randleslab.pratt.duke.edu