The Genetic History of the British Isles
Dabbling in ancient DNA. Learning a bit of my history. The modern Brits descend from Bell Beakers.
As an inveterate and incurable dilettante and geneticist, I have an interest in genetic approaches to understanding the deep history of humans. This discipline can go by a whole host of different names: paleogenomics, archaeogenetics, and genetic anthropology.1 Invariably, it involves the discovery and analysis of ancient or historical DNA.
There are a number of different sources I’ve turned to learn about this field, but some of the most detailed, accessible, and compelling writing comes from a Substack called Razib Khan's Unsupervised Learning. Additionally, I also really loved Who We Are and How We Got Here: Ancient DNA and the New Science of the Human Past by the eminent Harvard geneticist David Reich. I read Reich’s book a couple years after its publication as I was completing my PhD, and I’ve been reading Razib’s work for years now.2 I really encourage others to check both out.
Today, I’ll focus on the genetic history of the British Isles by way of a book I stumbled across called Saxons, Vikings, and Celts: The Genetic Roots of Britain and Ireland by Bryan Sykes.3 Why? Well, the 2006 book just happened to be included with my Audible membership, and I happen to have quite a bit of ancestry from the British Isles (~85%).4 Plus, Britain can be considered one of the important founding sites of modern genetics: evolution (Charles Darwin), population genetics (R. A. Fisher), and DNA (Watson & Crick). Because of Britain’s pivotal role in the history of biological science, it has built up invaluable resources and methods for studying their genetic history.5 This also includes an incredible sampling of ancient DNA, including nearly 600 genomes from pre-Roman conquest of Britain.

Saxons, Vikings, and Celts is an interesting work of science writing on ancient DNA from an important figure in the field. It attempts to marry historical narratives with genetic data. This is a heady and challenging venture made fraught by the sociopolitical importance certain historical narratives are attributed. And though it appears that many of Sykes’ claims have not aged well, he wasn’t all wrong either.
The limits on his accuracy are probably a function of his tools and sampling. In the pre-next-generation sequencing (NGS) era of $200 whole genomes, Sykes had to rely on haplogroup data from contemporary populations. In fact, all of his claims are premised on mitochondrial or Y chromosome data. This leaves out a lot of the genome, all of the non-sex chromosomes and the X chromosome, including all of the specific base information contained therein. We’ve learned a lot in genetics since 2006. Our tools have advanced, and we have a lot more samples of ancient DNA. Nonetheless, Sykes makes a great effort, and assembled an interesting book that is still worth reading.
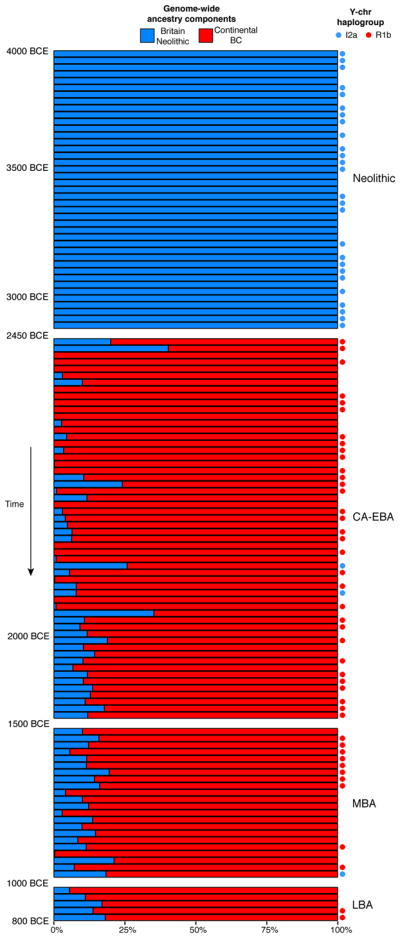
His central claim is that the modern residents of the British Isles are most similar to early European settlers of the Neolithic (12kya-6.5kya). Unfortunately, this central claim is likely incorrect. A 2018 Nature paper from the labs of David Reich and Carles Lalueza-Fox found that, between 2450-2000 BCE, over 90% of British DNA was replaced by European Steppe herders (the Bell Beaker culture) in a migration that brought significant amounts of Steppe DNA (i.e. the R1b haplogroup of the Y) to western and northern Europe. It is unclear exactly what led Sykes astray. It may have been a biased sample, the reliance on Y/mtDNA, or another issue with his analysis. Given that the Reich's lab paper presents its own case using Y haplogroup data, it seems that Sykes’ inaccuracy was more likely a function of his sample and analytical choices rather than the haplogroup method itself.
Brief Interlude on Haplogroup Analysis
Above I mentioned haplogroups, which may have thrown readers without backgrounds in genetics. I’ll try and explain.
Put simply, a haplotype is a group of alleles (alleles are different versions or variants of genes) in an organism that are inherited together from a parent. If you remember your high school biology class, including Mendel’s law of independent assortment and chromosomal recombination during meiosis, haplotypes are essentially the parts of chromosomes that resist this chopping and swapping - unbroken chunks. A haplogroup is a similar but extended version of this idea. They are groups of similar haplotypes that share a common ancestor that is marked by a single-nucleotide polymorphism (SNP).6 In human genetics, there are two types of commonly studied haplogroups; those of the Y chromosome and those of the mitochondrial genome (mtDNA).7
Haplogroup analysis is a method of studying chunks of DNA inherited together and how they change over time. The changes are tracked by comparing the genetic variations in certain regions and times, tracing them back to a common ancestor. A key to this analysis is that we know that less diverse distributions of haplogroups come from more recent populations. We also have good models of how frequently changes to a haplogroup are expected to occur. Ultimately, haplogroup analysis of ancient DNA samples helps researchers infer the geographic origin, migration patterns, and evolutionary relationships of ancient populations.
Before we circle back to Sykes’ book on Britain, let’t return to why relying solely on Y and mtDNA can be an issue when trying to learn about ancient population. To repeat myself, the simple answer is that it ignores a lot of the genome. The picture is necessarily incomplete. More specifically, the Y or mtDNA only reflects the separate paternal or maternal lineage, which may not even tell the same story.8 Additionally, haplogroups may be subject to genetic drift,9 selection,10 or admixture.11 This can distort the phylogenetic signal and slant the geographic distribution of haplogroups. Y or mtDNA may have low mutation rates or high homoplasy (convergent evolution), which can reduce the resolution or accuracy of the phylogenetic reconstruction too. There is the additional challenge of reconciling the Y and mitochondrial lineages. As mentioned above, they can tell different stories, especially because humans tend to have more female than male ancestors.12 However, I don't want to denigrate the utility of Y and mtDNA haplogroups. They are a very interesting and important source of ancestral genetic information. Sykes uses his haplogroup data to its fullest at least at the time of the publication of this book. Plus, he has a great understanding of history and culture, especially as a scientist.
Returning to Sykes’ Claims on UK Genetic Ancestry
Another big point that Sykes makes is that despite popular mythology, Anglo-Saxons make up a small portion of English ancestry. He pegs this number around but under 20% of the total ancestry, even in Southern England. A 2016 study in Nature Communication suggests the contribution from Anglo-Saxons to modern Brits is around one-third and in East Anglia approaches 40%.13 Sykes’ conclusion wasn’t quite on the money here but differs only modestly from the updated estimates.
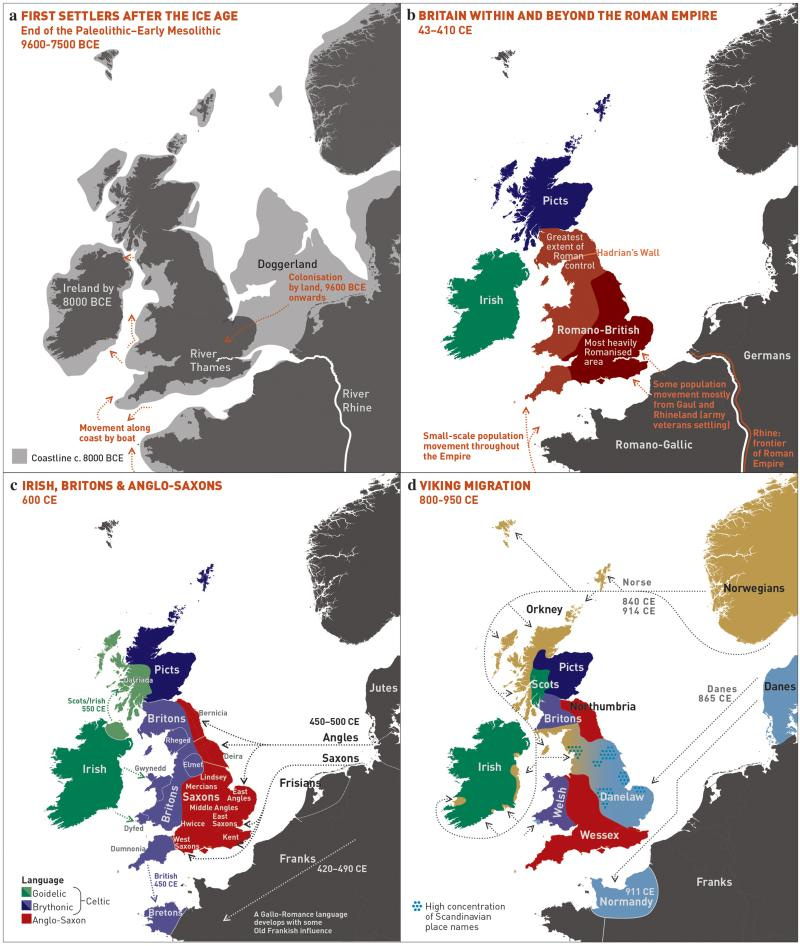
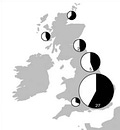
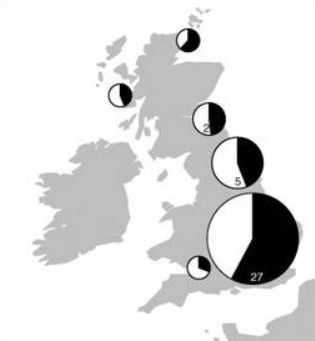
Let’s give Sykes more of his due though. He asserts that the Norman conquest (1066) only contributed 2% ancestry to current populations and that only traces of the Roman occupation remain. This observation still holds up. Further, he asserts the contribution from Vikings (Danes and Norwegians) was substantial but concentrated in central, northern and eastern England. He alleges a significant Viking contribution in the Orkney and Shetland Islands, in the vicinity of 40 percent, where the contribution was surprisingly balanced between maternal and paternal lines. Because otherwise the paternal and maternal lines suggest the outsized success of only a few male lines (not an unusual pattern in populations with histories of being subject to ancient conquests). I haven’t seen anything that contradicts this either.
Now, I haven't done the due diligence on running down everything that Sykes presents, but I think some of his claims and the overall project has plenty of merit even nearly two decades later. Regardless of the status of the claims, I appreciated learning more about the origins of my ancestry. The human species of course shares a relatively short and single genetic history - geneticists are often fond of saying we’re not that different from each other - but we’re also quite curious about how we do differ and who our nearest relatives in the past were. As my genome is in part descended from the seed of Albion, I appreciated that Sykes tried to disentangle these origins for me.
For the latest and best summary of the genetic origin of Brits, check out the writing of
:The genetic legacy of Ice-Age foragers in the ancestry of modern Britons is nearly nonexistent. Instead, the earliest settlers of the Isles were subsequently almost totally replaced by Neolithic farmers from the Mediterranean nearly 6,000 years ago. Then, 4,500 years ago, those farmers in turn saw themselves supplanted by a group saddled with the unwieldy label “Bell Beakers” (for the bell-shaped urns by which we first traced them). Genetically similar to contemporary Northern Europeans, the Bell Beaker migrants accounted for 90% of the ancestry of the British population by 2000 BC, with the Neolithic proportion dropping to 10% in five centuries.
He has a two-part series on this question:
Technically, these terms can refer to more specific domains, but for our purposes we can think of them as synonyms.
Before migrating to Substack, Razib ran a successful blog called Gene Expression.
This may have been a book that was recommended by Razib, which was probably why it caught my eye. Sykes was an important figure in the field of ancient DNA, who Razib regularly cites.
This comes with the usual caveat of direct-to-consumer ancestry testing. I’d love to write more about the technical issue involved here. For now, it is good to understand that these are estimates and not necessarily written in stone results. If you are a consumer of these services, you have probably seen revisions and updates to your ancestry scores.
The UK is still home to cutting edge research in genetics and genomics. The UK Biobank has been an enormously important resources and has supported numerous seminal findings in quantitative genetics.
A single nucleotide polymorphism (SNP) is a variation in a DNA sequence at a single position among individuals. It is a way of saying there is a change to a base of DNA at a given position that there are a certain portion of people in a population with and without that change.
This is because mtDNA is solely passed from mothers to children, and the Y is passed from fathers to sons.
Genetic drift is a change in the frequency of an existing gene variant (allele) in a population due to random chance.
Genetic selection is a process that changes allele frequencies in a population due to the fitness advantage afforded by those alleles.
Admixture is the presence of DNA in an individual from a distantly-related population or species.
The procreative success of men is distributed unequally, sometimes drastically so.